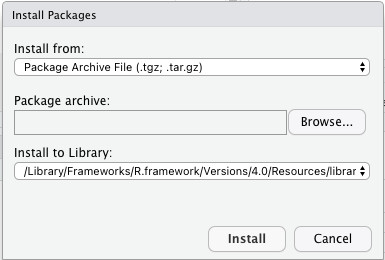
 I am able to install packages from R on the command-line. It also hangs if I try to install a new R package, showing the 'retrieving package installation context' message.  If I try to open a new R notebook document it hangs. To get a bug fix or to use a feature from the development version, you can install the development version of dplyr from GitHub. I installed Rstudio server on my ubuntu machine with R 3.5.1. Packages ‘devtools’, ‘lme4’ are not available (for R version 4.0. Installation The easiest way to get dplyr is to install the whole tidyverse: install.packages('tidyverse') Alternatively, install just dplyr: install.packages('dplyr') Development version. Installing packages into ‘C:/Users/ricardo.rfflj/Documents/R/win-library/3.4’ Please download and install the appropriate version of Rtools before proceeding: WARNING: Rtools is required to build R packages but is not currently installed. They all give the same error message, as shown below: > install.packages(c("devtools", "lme4")) Step 4: Select the language of your choice in the installer and click OK. Double click on the installer to launch it. Step 3: Clicking on the tab will download the R installer. Step 2: Click on the Download R 3.6.0 for Windows. I tried another command: install.packages (c ("devtools", "lme4")) Step 1: Go to the website CRAN R Project Windows. Everything went well until I tried to install a package using the command: install.packages ("ggplot2")  I installed version 4.0.2 of R for Windows 10 (圆4), then R tools 40 and finally RStudio. Install.packages("boot", repos = NULL, type = "source")īut I would prefer to do this with a single call to install.packages only and since install.packages is capable of downloading files anyway I feel this should be possible.I am taking a Data Science course at Coursera using R. We can work around this with the following two step procedure: download.file( Both actions open a window with fields for Variable. If you see RLIBSUSER, highlight it and click Edit. The Environmental Variables window pops up. Create a new virtual environment in a folder called myenv. Next I need to add this folder to the RLIBSUSER path: Click Start -> Control Panel -> User Accounts -> Change my environmental variables. Navigate into your RStudio project directory by using the following command: cd .The most common way is to use the CRAN repository, then you just need the name of the package and use the command install.packages. 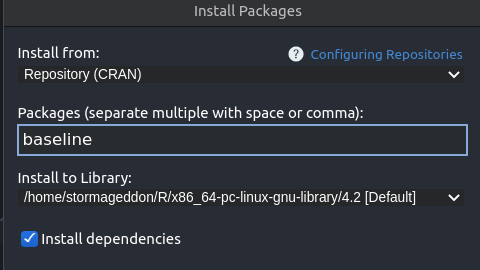 So, for publicly available packages, this means to what repository it belongs. How you can install an R package will depend on where it is located. It is recommended that you use one virtual environment per project, similar to how packrat is used to manage R packages within a project. How to Install an R Package Installing R Packages From CRAN. Suggesting that the URL is being interpreted as the package name, not its location. Step 2) Create a Python environment in your project. However this fails with: Warning in install.packages : Reading ?install.packages particularly the description of the pkgs argument suggests: install.packages( This is a similar question but it is different because it only describes how to install from local files not general URLs.įor the sake of this question I will use a link to the boot package source. I want to do this to make it easy for people to test a pre-release version of the package which should not be widely (or permanently) available. That will run some code in your R Console and will take a few seconds to install. I would like to install a package directly from a URL for the package source. When I tried to install tm, I got the following: > install.packages(tm) Warning in install.packages : unable to. In the Source Editor pane, type install.packages('readxl') and click Run to download the readxl library.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
